Researchers find that the methylation of a newly identified region of the INS-IGF2 gene is responsible for IGF2 expression in breast cancer tumors and in breast cancer cells.
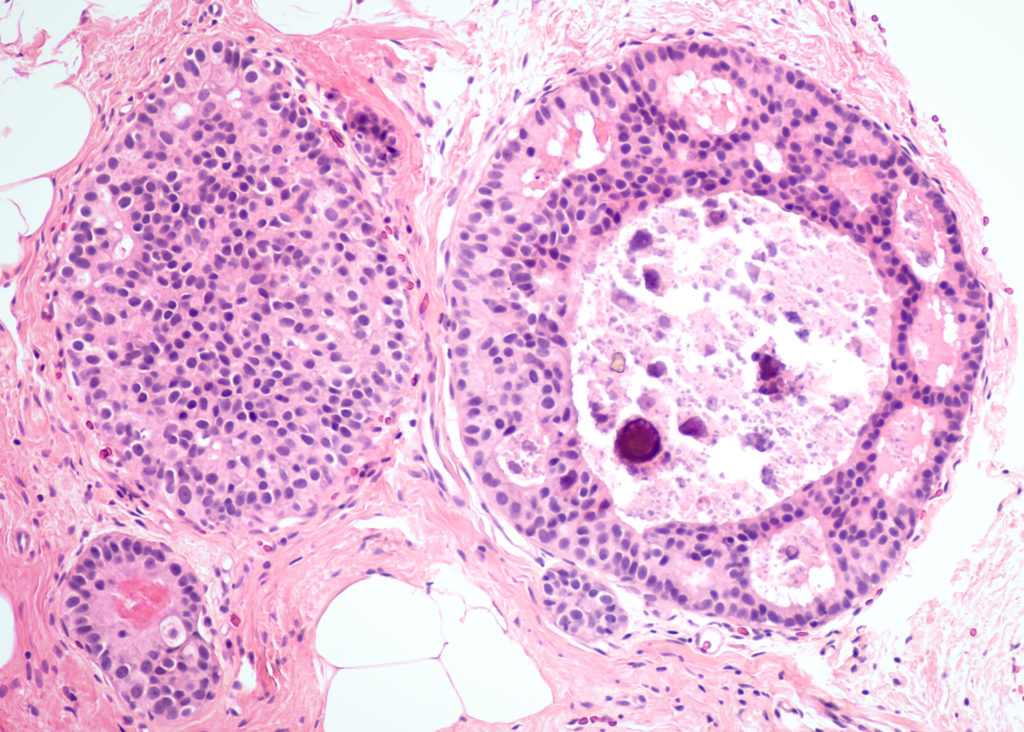
Determined to learn more and find new methods to eradicate breast cancer, a team of researchers turned their sights on a mechanism that could be a key in novel therapies: the IGF2 gene.
Based out of the United States’ California and Colorado, the researchers examined a particular gene in the DNA, located on Chromosome 11, and its functions in normal and breast cancer cells.
IGF2 Expression in Breast Cancer
The IGF2 gene is a growth factor that plays an important role in regulating stem cell activity and liver function in adults. This gene can be inhibited or stimulated by hormones, stress, nutrition, tumor suppressors, and oncogenes. The authors of this paper also note that IGF2 promotes cell proliferation, inhibits apoptosis, and stimulates the transformation of breast cancer cells.
Methylation in IGF2
Methyl groups are small molecules that are naturally introduced into the DNA at various times in human development—for a number of reasons. In short, the function of the methyl group is to ensure processes within the DNA operate the way they are biologically intended to. Methylation can change the activity in a segment of DNA without changing the sequence, and, when located in a gene promoter, can repress gene transcription.
In this study, the researchers note that the IGF2 gene is transcriptionally regulated through the methylation of five different promoters and DMRs (differentially methylated DNA regions).
“Abnormal methylation of IGF2 leads to various metabolic disorders like breast cancer, pancreatic cancer, diabetes, and endocrine related disorders [24–26].”
Researchers focused on analyzing the IGF2 gene and its DNA methylation patterns in normal and tumor tissue cell lines, which they obtained from breast cancer patients among four different ethnic groups.
New Region of INS-IGF2 Gene
In their studies, the researchers found a particular region within the INS-IGF2 locus, named INS-IGF2 DVDMR, that had anomalous methylation. Surprisingly, the hypomethylation (or loss of a methyl group) closely correlated with an increase in IGF2 mRNA and protein levels in all four breast cancer cell lines.
“We identified a new region with variable methylation within the INS-IGF2 locus, between exons 3 and 4, which we named INS-IGF2 DVDMR.”
The team demonstrated that when IGF2 levels are increased or decreased, they may inhibit or stimulate mitochondrial proteins in breast cancer cell lines, which in turn prevents cell death and produces chemoresistance.
“Since mitochondria are important targets of chemotherapy, IGF2 expression by breast cancer tumors may confer mitochondrial protection, thereby, inducing chemoresistance and effectively reducing clinical treatment efficacy.”
Conclusion
“IGF2 is highly expressed in breast cancer patients and ‘free’ circulating IGF2 levels in humans are significantly correlated with breast tumor size [8].”
Given that mitochondrial proteins were found to be significantly correlated with IGF2 levels in tissues from breast cancer patients, the researchers proposed that IGF2 represents a potential therapeutic target to decrease chemoresistance and improve survival among breast cancer patients.
“Our data also showed that the methylation profile of the INS-IGF2 DVDMR in paired breast tissues was distinct and could differentiate malignant breast tissue (hypomethylated) from normal adjacent breast tissue, which was hypermethylated.”
The results from this study suggest that methylation of the INS-IGF2 DVDMR is a key regulator of IGF2 expression in breast cancer.
—
Oncotarget is a unique platform designed to house scientific studies in a journal format that is available for anyone to read – without a paywall making access more difficult. This means information that has the potential to benefit our societies from the inside out can be shared with friends, neighbors, colleagues and other researchers, far and wide.
Click here to read the full scientific paper, published in Oncotarget.