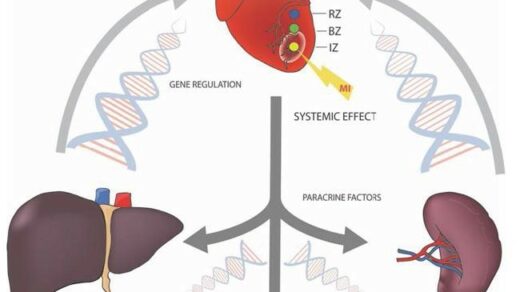
The increased expression of DLST has recently been associated with increased tumor aggression and a poor prognosis in neuroblastoma and triple-negative breast cancer.
The Trending With Impact series highlights Oncotarget publications attracting higher visibility among readers around the world online, in the news, and on social media—beyond normal readership levels. Look for future science news about the latest trending publications here, and at Oncotarget.com.
—
Dihydrolipoamide S-succinyltransferase (DLST) is a crucial gene/protein/enzyme involved in the oxidative phosphorylation (OXPHOS) pathway and cellular energy production. Recent studies have demonstrated that, in neuroblastoma and triple-negative breast cancer (TNBC), increased expression of DLST is associated with increased tumor aggression and a poor disease prognosis. Researchers also found that, in leukemia and TNBC cell lines, the knockdown of DLST leads to apoptosis. These findings suggest that neuroblastoma and TNBC may benefit from DLST-inhibiting cancer therapy.
In light of this evidence, researchers Christina Kuhn, Myriam Boeschen, Manuel Philip, Torsten Schöneberg, Doreen Thor, and Susanne Horn from the University of Leipzig, University Duisburg-Essen and the German Cancer Consortium investigated approved drugs that may target DLST-activated tumors. In their recent study, the team used data from the Genomics of Drug Sensitivity in Cancer (GDSC) project to identify new drug candidates for the treatment of DLST-activated tumors. On January 12, 2023, their research paper was published in Oncotarget’s Volume 14, entitled, “Candidate drugs associated with sensitivity of cancer cell lines with DLST amplification or high mRNA levels.”
The Study
“With the advent of complex genetic datasets of roughly 1000 cell lines in the Cancer Cell Line Encyclopedia (CCLE) and on drug resistance in the Genomics of Drug Sensitivity in Cancer project (GDSC), analyses of drug sensitivity have become possible on a larger scale [6, 7].”
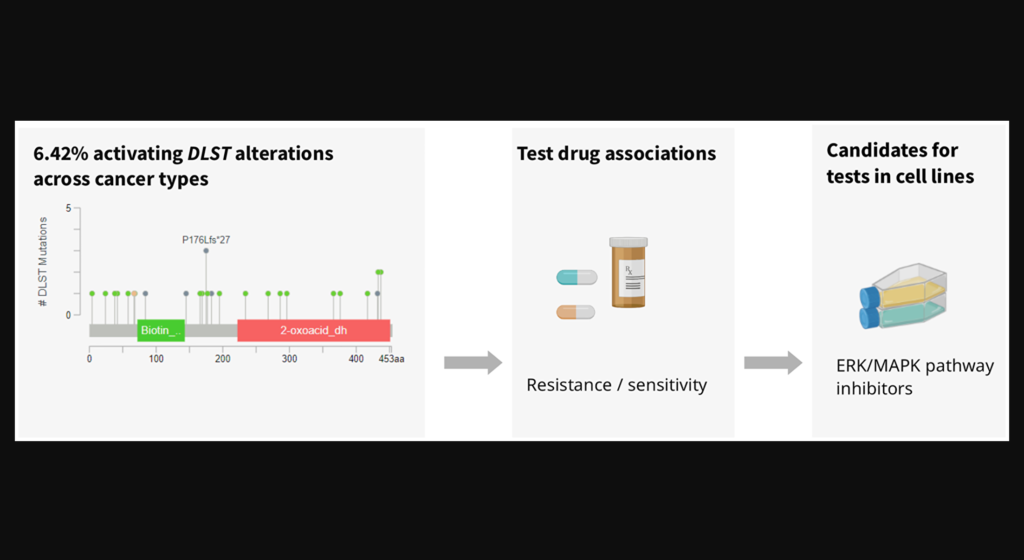
Since neuroblastoma and TNBC tumor cell viability may be DLST-dependent, DLST is a promising target for cancer therapy. The researchers used the Cancer Cell Line Encyclopedia (CCLE) to identify cancer cell lines with DLST amplifications or high mRNA levels. They then measured the sensitivity of these DLST+ cell lines to 250 drugs in the GDSC dataset and compared the data to a subset of cell lines without DLST amplifications or high mRNA levels.
“To identify drugs that inhibit viability of specifically DLST-activated tumor cells, we compared cell lines with supposedly activating changes of DLST (DNA amplification, high mRNA levels) to cell lines without DLST changes.”
Results & Conclusions
“DLST-altered cell lines were more sensitive to 7 approved drugs, among these obatoclax mesylate, a BCL2 inhibitor that reduces OXPHOS in human leukemia stem cells.”
The researchers identified seven drug candidates that demonstrated significantly higher sensitivity in DLST+ cell lines than in the control cell lines. In addition to a BCL2 inhibitor found to reduce OXPHOS, multiple protein kinase inhibitors were identified as efficient in the DLST+ cell lines. This suggests that DLST-altered cell lines may also be vulnerable to ERK/MAPK pathway-targeting drugs. The researchers propose that the drug candidates identified in this study warrant further drug efficacy testing in knock-in cell lines and DLST-activated tumors.
“We therefore conclude that, in addition to OXPHOS, protein kinases could be potential targets of therapy in the presence of DLST amplifications or high mRNA levels.”
Click here to read the full research paper published by Oncotarget.
ONCOTARGET VIDEOS: YouTube | LabTube | Oncotarget.com
—
Oncotarget is an open-access, peer reviewed journal that has published primarily oncology-focused research papers since 2010. These papers are available to readers (at no cost and free of subscription barriers) in a continuous publishing format at Oncotarget.com. Oncotarget is indexed/archived on MEDLINE / PMC / PubMed.
For media inquiries, please contact media@impactjournals.com.