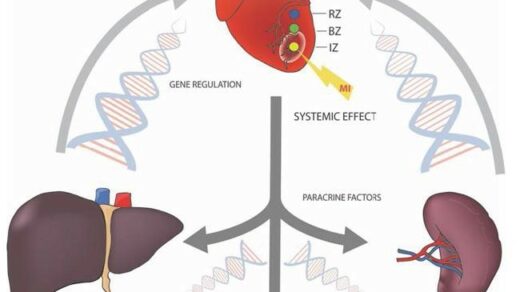
Researchers determined the prognostic ability of three NOTCH1 gene variants by incorporating them into two non-tumorigenic breast cell lines.
The Trending With Impact series highlights Oncotarget publications attracting higher visibility among readers around the world online, in the news, and on social media—beyond normal readership levels. Look for future science news about the latest trending publications here, and at Oncotarget.com.
—
The genetic changes that occur within the protein-coding gene NOTCH1 have not yet been fully studied or classified. Despite a lack in research, previous studies have suggested that NOTCH1 may be a potential target for novel cancer therapies, particularly against triple-negative breast cancer (TNBC). NOTCH1 variants in TNBC tend to cluster in the PEST region and have previously been linked to gamma secretase inhibitor (GSI) sensitivity and chemotherapy resistance.
“Furthermore, TNBC patients with increased Notch1 expression have demonstrated increased aggressive phenotypes and lower median overall survival [25].”
Since TNBC is well-known for a lack of actionable therapeutic targets, aggressive phenotypes and poor prognoses, there is an important need to develop new targeted therapies—as well as predictive markers for those therapies. Researchers from The Johns Hopkins University School of Medicine, Vanderbilt University Medical Center and The Vanderbilt-Ingram Cancer Center experimented in vitro with NOTCH1 variants and their ability to predict TNBC responsiveness to GSIs and standard of care chemotherapies. Their trending research paper was published by Oncotarget on February 16, 2022, and entitled, “NOTCH1 PEST domain variants are responsive to standard of care treatments despite distinct transformative properties in a breast cancer model.”
The Study
The researchers used three publicly available tumor-associated variant databases to identify three NOTCH1 variants that are commonly mutated in breast cancers; two variants were located in the A2441 site on NOTCH1 and the third variant was located in the PEST region of NOTCH1. To investigate the role of these NOTCH1 variants in TNBC in vitro, the team cultured two non-tumorigenic breast epithelial cell lines. Uniquely, they used an adeno-associated virus (AAV) vector to isogenically incorporate the NOTCH1 variants into the two cell lines. The researchers also developed a wildtype vector for the control arm of the study.
“In addition to the NOTCH1 variants, a targeted wildtype (TWT), which underwent the same gene targeting mechanism with a wildtype vector, was generated for both parental cell lines to act as a control.”
A standard growth factor supplemented media was used to determine if the NOTCH1 variants caused increased proliferation in the non-tumorigenic cell lines. Compared to the controls, no significant change in proliferation was observed. They also removed the epidermal growth factor (EGF) from the cells to determine if these NOTCH1 variants impart a ligand-independent proliferative advantage. In both cell lines, their results demonstrated that the A2441 variants exhibited EGF-independent growth, while the PEST NOTCH1 variant did not. Immunoblot analyses suggested that, in the absence of EGF, the A2441 NOTCH1 variants activated the MAPK pathway. These EGF-independent NOTCH1 variants (not the PEST NOTCH1 variant) conferred an invasive growth phenotype, increased migratory potential, had dysregulated 3D morphology, and significantly altered gene expression in cancer pathway genes.
Next, to measure the responsiveness and susceptibility of these variants to therapeutic agents, the cells were treated with six chemotherapeutic agents and nirogacestat—a GSI drug. Interestingly, none of the three variants demonstrated significantly different responses to the treatments when compared to one another. Furthermore, all of the variants showed sensitivity to these standard therapies used against TNBC. This suggests that these specific genetic changes within NOTCH do not have a large impact on tumor behavior and may not be useful as predictive markers for therapy response.
Conclusion
“Taken together, these data suggest that the oncogenic potential of NOTCH1 PEST domain variants depends on both variant type and amino acid location.”
Contrary to previous studies, the researchers found that the three NOTCH variants did not demonstrate significantly different responses to the GSI or the chemotherapies despite demonstrating distinct phenotypes. The lack of differential responses demonstrated by the variants in this study suggests that there is high variability among NOTCH1 variants. The prognostic potential of NOTCH1 may be dependent on the type of variant and its location, but more expansive research is necessary.
“Future studies involving meticulous characterization of an expansive panel of NOTCH1 variants in a similar model may provide mechanistic insight and predictive and/or prognostic value that is both variant type and site dependent.”
Click here to read the full research paper published by Oncotarget.
ONCOTARGET VIDEOS: YouTube | LabTube | Oncotarget.com
—
Oncotarget is a unique platform designed to house scientific studies in a journal format that is available for anyone to read without a paywall making access more difficult. This means information that has the potential to benefit our societies from the inside out can be shared with friends, neighbors, colleagues, and other researchers, far and wide.
For media inquiries, please contact media@impactjournals.com.