In the cover paper chosen for Oncotarget’s Volume 12, Issue 16, researchers from Yale University stratified subtypes of a rare and aggressive form of cancer: leiomyosarcoma (LMS).
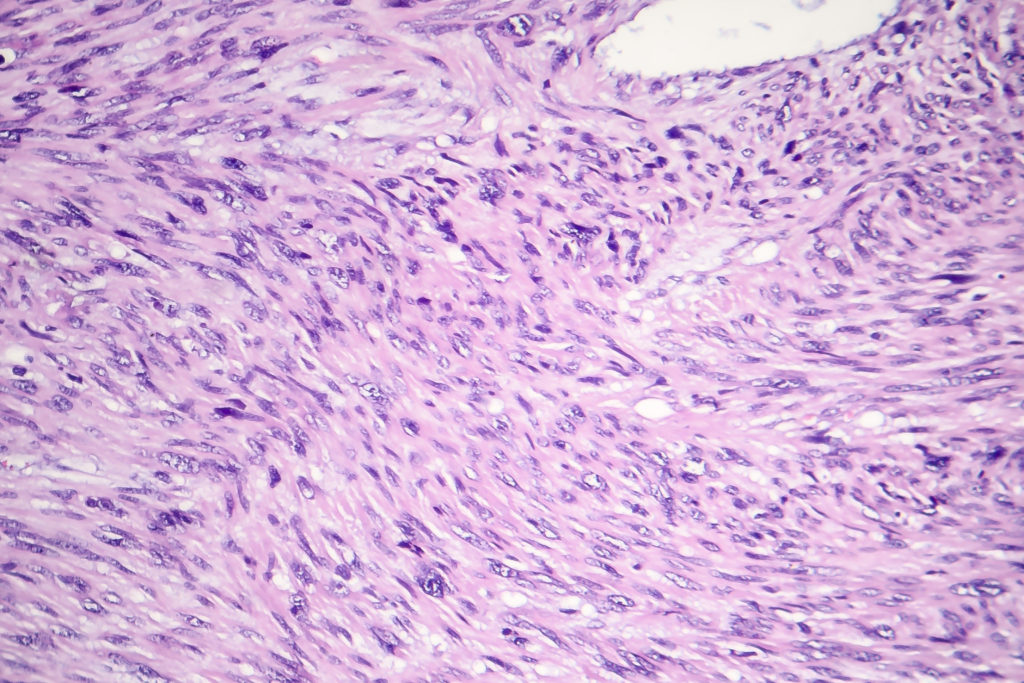
The human body consists of muscles that are either voluntarily or involuntarily maneuvered—meaning the brain does not have conscious control over involuntary muscles. Smooth muscles, such as those present in the blood vessels, gallbladder, intestines, stomach walls, urinary system, uterus, prostate, and so on, are involuntary muscles. Diverse mesenchymal soft tissue sarcomas can form in involuntary muscles. These rare and aggressive sarcomas are known as leiomyosarcomas (LMS).
“Newly diagnosed patients are at high risk of distant recurrence and poor disease-specific survival [2].”
While researchers find that alterations in the epigenome are often what initiates aspects of cancer, epigenetic profiling of LMS has been limited. In 2021, researchers from Yale University School of Medicine conducted a new study in search of specific differences in LMS subtypes using epigenetic profiling and integrated analysis. The goal of their study was to identify potential targets for therapeutic intervention and diagnosis. This paper was chosen as the cover paper published in Oncotarget’s Volume 12, Issue 16, and entitled, “Epigenetic signatures differentiate uterine and soft tissue leiomyosarcoma.”
“Only a limited number of studies have compared the outcomes based on LMS subtypes despite obvious clinical differences.”
The Study
“In this study, we performed a comprehensive analysis and compared the DNA methylation and RNA expression profiles of ULMS and STLMS samples from the TCGA-SARC study.”
First, the team identified two LMS subtypes based on their site of origin: uterine LMS (ULMS) and non-uterine soft tissue LMS (STLMS). Next, they compared their molecular landscapes using The Cancer Genome Atlas-Sarcoma (TCGA-SARC) dataset and comprehensively analyzed 27 ULMS and 53 STLMS samples. Researchers assessed for epigenetic and transcriptomic differences between ULMS and STLMS. They hoped these differences would help identify differentially methylated genes and differentially expressed genes (DEGs) in these LMS subtypes. Findings from the TCGA-SARC dataset were then compared to two independent DNA methylation datasets, GSE140686 and GSE68312.
“In this study, we performed an integrated analysis of 98 clinically derived LMS samples and 11 controls using three publicly available datasets to identify epigenetic changes which characterize LMS subtypes and may be used as clinical biomarkers and therapeutic targets.”
Datasets were then used in comparative DNA methylation analysis to compare controls with the identified epigenetic changes associated with tumorigenesis. Network and pathway analysis were performed to further characterize the differential transcription signatures between ULMS and STLMS.
Results and Conclusion
“Principal Component Analysis (PCA) of the DNA methylation data revealed separation between ULMS and STLMS.”
The researchers found that the majority of the STLMS and ULMS samples segregated themselves and formed spatially distinct clusters. ULMS and STLMS tumors were found to be epigenetically distinct subtypes. 283 differentially methylated regions in promoter-associated CpG island sites were identified that differ between ULMS and STLMS. Interestingly, upstream regulator analysis of the differentially methylated regions using Ingenuity Pathway Analysis revealed that this differential expression involves chromatin-modifying enzymes, DNA-binding proteins, and chromatin-binding proteins. This suggests that chromatin modulation may be involved in the oncogenesis of LMS subtypes.
“In summary, we show evidence for differential DNA methylation profiles between ULMS and STLMS suggesting that differential epigenetic profiles are associated with LMS subtypes and may be responsible for differences seen in clinical outcomes. These findings suggest that epigenetic profiles can be used to stratify patients and apply therapeutic agents targeting the epigenetic mechanisms for LMS management.”
Click here to read the full research paper, published by Oncotarget.
YOU MAY ALSO LIKE: More Oncotarget Videos on LabTube
—
Oncotarget is a unique platform designed to house scientific studies in a journal format that is available for anyone to read—without a paywall making access more difficult. This means information that has the potential to benefit our societies from the inside out can be shared with friends, neighbors, colleagues, and other researchers, far and wide.
For media inquiries, please contact media@impactjournals.com.