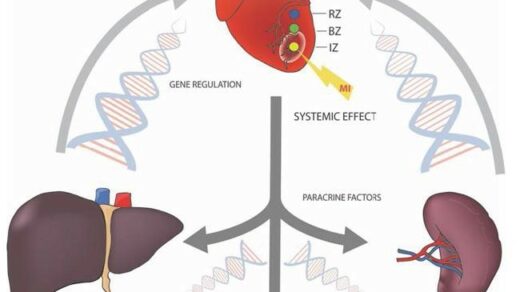
Researchers used the CRISPR-Cas9 tool to analyze a p53 mutation in triple-negative breast cancer.

Research reports that genetic mutations arise within the p53 gene in 70% of all human cancers. This unstable protein is an important transcription factor and tumor suppressor with additional transcription-independent functions, including a role in the regulation of DNA replication. Forms of the mutated p53 (mtp53) gene found in cancer are often highly stable proteins that can gain new functions—such as R273H. Mutant p53 R273H proteins gain functional properties which promote oncogenesis in triple-negative breast cancer, in part by modulating the cell’s replication machinery.
In a 2021 study, researchers from the City University of New York, Columbia University, and Weill Cornell Medical College further investigated mutant p53 R273H in triple-negative breast cancer. The team used CRISPR-Cas9 targeting of the mutant p53 gene in triple-negative breast cancer cell lines to further understand the effects of reducing mutant p53 proteins through artificial RNA (shRNA) mediation. Their paper was chosen as the cover of Oncotarget’s Volume 12, Issue 12, and entitled, “Frame-shift mediated reduction of gain-of-function p53 R273H and deletion of the R273H C-terminus in breast cancer cells result in replication-stress sensitivity.”
“In this research perspective we used CRISPR-Cas9 targeting of these C-terminal regions of the mtp53 gene in MDA-MB-468 [breast cancer] cells to further extend our findings with shRNA-mediated reduction of mtp53 proteins [18–20].”
CRISPR & CRISPR-Cas9
In 1987, researchers from Japan first published about clustered regularly inter-spaced short palindromic repeats in the DNA sequence. In 2002, a team of Dutch scientists shortened the name of this prevalent pattern to CRISPR. Within the year, the same Dutch team noticed a cluster of genes that always accompanies the CRISPR sequence—which they named CRISPR-associated genes, or Cas. Introduced in 2007, CRISPR-Cas9 is a technique that uses relatively new technology, and is now renowned in the scientific community for its simplicity and accuracy in gene editing ability.
CRISPR-Cas9 enables researchers to edit the DNA sequence by removing, adding, or altering specific sections of the genome. CRISPR-associated protein 9 (Cas9) is an enzyme, capable of severing both strands of DNA. The enzyme can cut DNA at a specific location in the genome, with localizing help from a guide RNA (gRNA) sequence. Once the DNA has been cut, the cell recognizes that the DNA is damaged and begins to repair itself using DNA repair machinery. DNA repair machinery can be used to introduce changes in the genome.
The Study
The researchers in this study wrote that both the C-terminal domain (CTD) and oligomerization domain (OD) of mtp53 R273H proteins are intact, and it is not clear if these regions are responsible for chromatin-based DNA replication activities. They used the CRISPR-Cas9 tool to edit the C-terminal regions of mtp53 R273H in breast cancer cells to examine their ability to influence cell proliferation, DNA replication, and cell cycle progression.
“CRISPR-Cas9 sgRNA editing of the C-terminal regions of the endogenous mtp53 gene were carried out so as to delete gene sequences that correspond to the OD and CTD regions.”
The team generated breast cancer cells and edited CTD and OD regions of mtp53 R273H using the CRISPR-Cas9 tool. They then treated the cell populations with thymidine—to block cells at G1/S phase in the cell cycle. The researchers then compared the status and proliferation of the variants with the original cell line and observed changes in total cell lysates by western blot analysis.
“We examined how changes in the level of mtp53 R273H level and/or deletion of the CTD, or OD plus CTD, region influenced cell proliferation, cell cycle progression, and chromatin association of mtp53, RPA, PCNA and MCM2.”
Results
Having deleted or frame-shifted gene sequences corresponding to the OD and CTD regions of mtp53 R273H, the researchers found that mtp53 R273H CRISPR-Cas9 variants proliferated more slowly than the parental cells, and the frameshift variant mtp53 R273Hfs387 expressed the most extreme low levels of mtp53. They also reported a number of other important findings.
“We observed reduced mtp53 protein level for mtp53 R273Hfs387 cells, and reduced proliferation in the associated CRISPR-Cas9 clone (see Figures 1 and 2).”
Conclusion
“Although currently we are unable to articulate the precise roles of these distinct regions of p53 in response to thymidine, our studies suggest that they may function at temporally distinct stages of S-phase.”
Click here to read the full paper, published by Oncotarget.
WATCH: MORE AGING VIDEOS ON LABTUBE TV
- Immune Checkpoint Inhibitors May Improve Outcomes in High-Risk Solid Tumors
- Mapping the Hidden Structure of Glioma Research: What Are We Missing?
- SCD1 Inhibition Strategy Shows Potent Synergy with Regorafenib and Metformin in Tumor Cell Killing
- Oncotarget Editorial Highlights Advances in Scientific Integrity and Publishing Transparency
- CREB5: A Master Regulator of Stem Cell-Like Programs in Prostate Cancer Progression
—
Oncotarget is a unique platform designed to house scientific studies in a journal format that is available for anyone to read—without a paywall making access more difficult. This means information that has the potential to benefit our societies from the inside out can be shared with friends, neighbors, colleagues, and other researchers, far and wide.
For media inquiries, please contact media@impactjournals.com.